


|
|
Home |
DynOmics 1.0 |
Tutorials |
Theory |
References |
iGNM 2.0 |
Mol. Sizer&Timer |
ANM 2.0 |
Comp. Biol. Lab |
PITT site
|
Theory
Environment GNM (envGNM)
Gaussian Network Model (GNM) is a powerful tool to sample conformational dynamics based on contact topology in a coarse-grained presentation. Here we present a method to consider protein dynamics in the presence of ‘environment’ (1). The ‘environment’ here can be crystal contacts defined in structures solved by x-ray crystallography, a protein in homo-/hetero-dimers, a part of a protein complex, the DNA in complexed with transcription factors, or even a large ligand (or ligands) in a protein. The protein dynamics, in the presence of these ‘environments’, can be assessed in a more rigorous physics ground (the size and number of the orthogonal normal modes are the same as those in unperturbed systems (in the absence of the environment)) with our theories (2) that guarantee our implementations to be more efficient (saving 3/4 time, assuming equal size of system and environment), more memory friendly (saving >1/2 time) and more accurate (enhanced correlation between predictions and B-factors). The environment consideration in ANM theory was published (1) and reviewed (2); its GNM counterpart is first derived in this work and proven to be more accurate in the B-factor predictions than conventional GNM.
envGNM can be derived as follows.
Start from potential energy function of GNM,
![]() (1)
(1)
![]() (2)
(2)
In a
N node system, the state vector is ![]() and
the state vector represent the system difference between instantaneous state
and equilibrium state;
and
the state vector represent the system difference between instantaneous state
and equilibrium state; ![]() are
a Hessian matrix of GNM model and a spring constant, respectively.
are
a Hessian matrix of GNM model and a spring constant, respectively.
![]() is
a N by N topology matrix, which the off-diagonal element
is
a N by N topology matrix, which the off-diagonal element ![]() =
-1 if node I is contact with node j; the diagonal element
=
-1 if node I is contact with node j; the diagonal element ![]() .
.
Separate
the total system to system and environment, so ![]() can
be represent by 4 sub blocks:
can
be represent by 4 sub blocks: ![]() ,
,![]() ,
,![]() ,
,![]()
![]() (3)
(3)
![]() (4)
(4)
![]() contains
the Hessian elements of the protein/DNA system;
contains
the Hessian elements of the protein/DNA system;![]() have
only Hessian elements of the environment;
have
only Hessian elements of the environment; ![]() and
and
![]() are
the Hessian elements that represent the interactions between the protein/DNA
system and the environment.
are
the Hessian elements that represent the interactions between the protein/DNA
system and the environment.
On the other hand, ![]() can
be split into 2 parts,
can
be split into 2 parts, ![]() and
and
![]() where
n is the number of nodes of the system, and rewrite to
where
n is the number of nodes of the system, and rewrite to
![]() (5)
(5)
Here ![]() and
and
![]() are
the displacement vectors of the protein/DNA system and the environment,
respectively.
are
the displacement vectors of the protein/DNA system and the environment,
respectively.
Now, we turn the form of Eq. 2 into system and environment representation.
![]() (6)
(6)
The small perturbations from environment will not affect the
system when the system is at equilibrium, ![]() .
Therefore, Eq. 6 can be simplified at this condition.
.
Therefore, Eq. 6 can be simplified at this condition.
![]() (7)
(7)
![]() (8)
(8)
Substitution of Eq. 8 into Eq. 6 we obtain
![]() (9)
(9)
![]() (10)
(10)
![]() (11)
(11)
Consider the middle term of Eq. 11 as a new Hessian,![]() :
:
We
define ![]() (12)
(12)
to have
![]() (13)
(13)
From Eq. 1 and Eq. 2
![]() (14)
(14)
Where the ![]() is a Kronecker product operator
which has the following properties:
is a Kronecker product operator
which has the following properties:
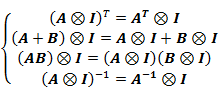
![]() can
be separated into 4 blocks similar to Eq. 3,
can
be separated into 4 blocks similar to Eq. 3,
![]() (15)
(15)
Substitute Eq. 15 into Eq. 14 to get
![]() (16)
(16)
Therefore, ![]() can
be rewrite as
can
be rewrite as
![]() (17)
(17)
Simplify Eq. 17 base on the properties of Kronecker product operator,
![]() (18)
(18)
![]() (19)
(19)
![]() (20)
(20)
Consider the first part of Eq. 20 as a new topology
matrix,![]()
![]() (21)
(21)
Substitute Eq. 21 into Eq. 13to get
![]() (22)
(22)
So that we can obtain the covariance:
![]() (23)
(23)
Environment ANM (envANM)
Anisotropic Network Model (ANM) is another powerful and widely used network model. The theory of envANM can be similarly derived as envGNM (above) (1,2).
Reference
1. Ming, D. and Wall, M.E. (2005) Allostery in a coarse-grained model of protein dynamics. Phys. Rev. Lett., 95, 198103.
2. Bahar, I., Lezon, T.R., Bakan, A. and Shrivastava, I.H. (2010) Normal Mode Analysis of Biomolecular Structures: Functional Mechanisms of Membrane Proteins. Chem. Rev., 110, 1463-1497.